Full Reviewed Journal Articles
Cao B, Yu X, Gonzalez G, Murthy AR, Li T, Shen Y, Yao S, Wang X. CGPA: a multi-context cancer gene prognosis atlas. Mol Cancer Res. 2026 Jan 13. doi: 10.1158/1541-7786.MCR-24-1186. Epub ahead of print. PMID: 41528384.
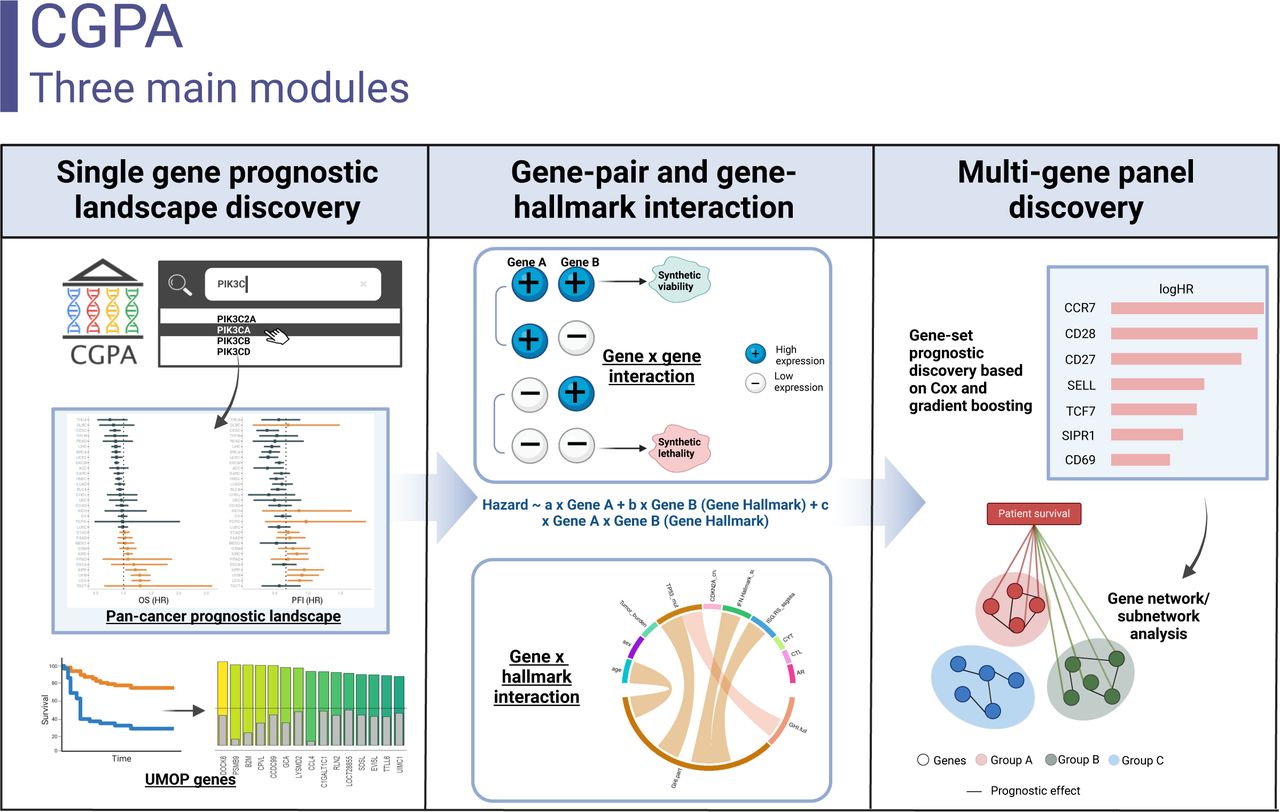
Natarajan U, Rathinavelu A. Regulation of PD-L1 Expression by SAHA-Mediated Histone Deacetylase Inhibition in Lung Cancer Cells. Cancers (Basel). 2025;17(17):2919. Published 2025 Sep 5.
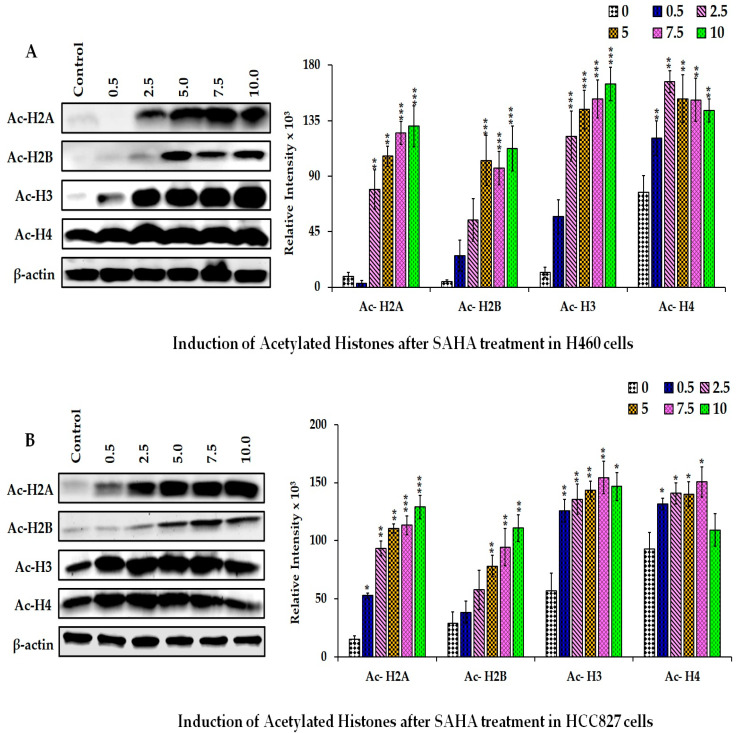
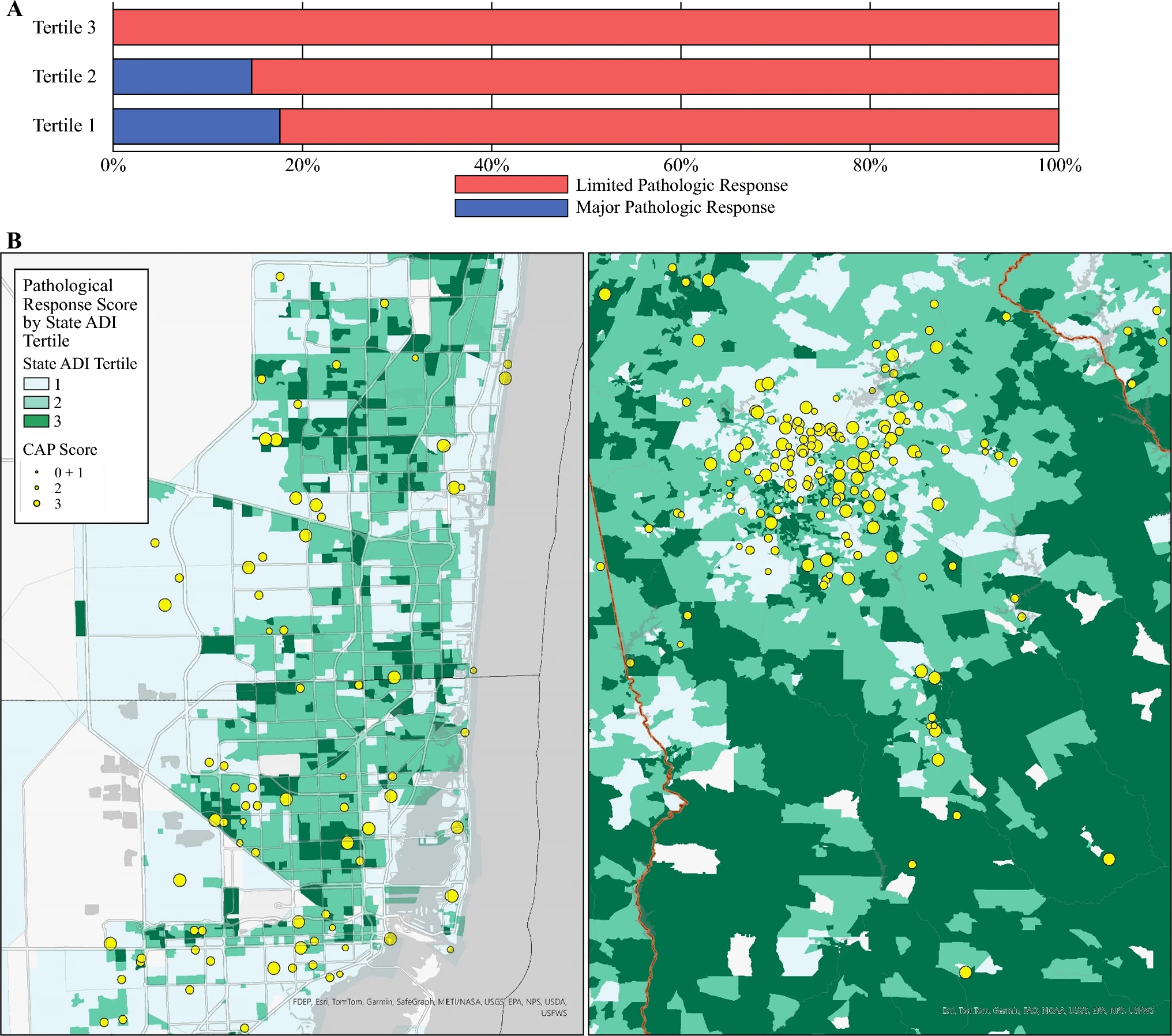
Martos, M. P.; Lee, M. S.; Merchant, N. B.; Kobetz, E.; Hester, C. A.; Stathias, V.; Maithel, S. K.; Schürer, S.; Datta, J. Neighborhood Disadvantage is Associated with Worse Pathologic Response in Pancreatic Cancer. Annals of Surgical Oncology 2025.
Schulz, J. M., Schürer, S. I., Reynolds, R. C., & Schürer, S. C. (2025). PRosettaC outperforms AlphaFold3 for modeling PROTAC ternary complexes. Scientific reports, 15(1), 37620.

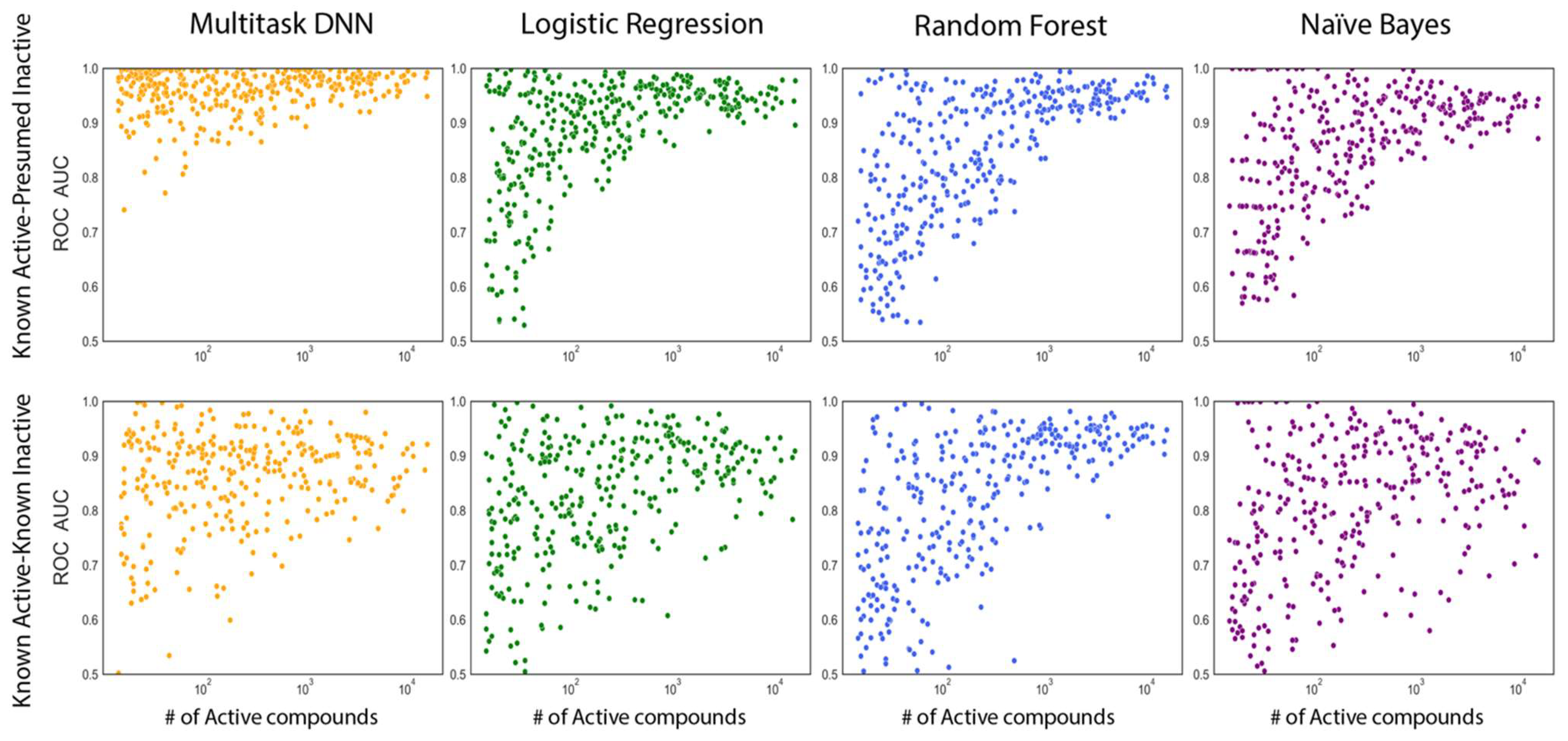
Hu, J.; Allen, B. K.; Stathias, V.; Ayad, N. G.; Schurer, S. C. Kinome-Wide Virtual Screening by Multi-Task Deep Learning. Int J Mol Sci 2024, 25 (5).
Full Article Preprints
Schulz, J. M.; Schürer, S. I.; Reynolds, R. C.; Schürer, S. C. Benchmarking the Builders: A Comparative Analysis of PRosettaC and AlphaFold3 for Predicting PROTAC Ternary Complexes. Research Square 2025, rs.3.rs-6866610;
DOI: 10.21203/rs.3.rs-6866610/v1.
Sobhan, M.; Islam, M.M.; Mondal, A.M. Interpreting Lung Cancer Health Disparity at Transcriptome Level. bioRxiv2025 2025;
DOI: 10.1101/2025.01.09.632292.
Ocasio, B. A.; Hu, J.; Stathias, V.; Martinez, M. J.; Burnstein, K. L.; Schurer, S. C. Pan-Cancer Drug Sensitivity Prediction from Gene Expression using Deep Learning. bioRxiv 2024.
Abstracts
Sinclair M.; Obregon C.; Chung C.; Vidovic D.; Rupprecht L.; Pissinis J.; Pilarczyk M.; Sotolongo F.; Jagodnik K. M.; Krenz T.; Natarajan U.; Kobetz E.; Mondal A. M.; Cen L.; Wang X.; Rathinavelu A.; Chalk S.;Bian J.; Lee J. H.; Song Q.; Stathias V.; Schürer S. C. Abstract 1086: Introducing the Florida Cancer Research (FL CARES) Network and the Platform for Accelerating Collaborative Computational Cancer Research (PAC3R). Cancer Research 2025, 85(8_Supplement_1), 1086–1086;
DOI: 10.1158/1538-7445.AM2025-1086.
Pissinis J. I.; Mazariegos O.; Pilarczyk M.; Sinclair M. S.; Rupprecht L.; Chung C.; Sotolongo F.; Lichtenstein G.; Jagodnik K. M.; Figueredo J.; Seo P.; Obregon C. J.; Blicharz M.; Smol P.; Pyrkosz M.; Jura M.; Stathias V.; Schürer S. C. Abstract 1081: Sylvester Data Portal: Streamlined access to standardized clinicogenomic data. Cancer Research 2025, 85(8_Supplement_1), 1081–1081;
DOI: 10.1158/1538-7445.AM2025-1081.
Mendel O.; Sinclair M. S.; Jagodnik K. M.; Krenz T.; Pissinis J.; Stathias V.; Bartal A.; Schürer S. C. Abstract1052: Integrating geographical and socioeconomic data with genomic information for enhanced prediction of cancer risk using large language models and knowledge graphs. Cancer Research 2025, 85(8_Supplement_1), 1052–1052;
DOI: 10.1158/1538-7445.AM2025-1052.
Alssamani F. H.; Natarajan U.; Jaganathan S.; Rathinavelu A. Abstract 223: Evaluating the therapeutic potential of MDM2 inhibitor RG-7388 and epigenetic modulator CM-272 in cisplatin-resistant ovarian cancer. Cancer Research 2025, 85(8_Supplement_1), 223–223;
DOI: 10.1158/1538-7445.AM2025-223.
Tumati N.; Natarajan U.; Jaganathan S.; Rathinavelu A. Abstract 441: Evaluation of the ehects of MDM2 inhibitor and epigenetic modifiers in combination with AURKB inhibitors for treating lung cancer. Cancer Research 2025, 85(8_Supplement_1), 441–441;
DOI: 10.1158/1538-7445.AM2025-441.
Jagarlamudi A.; Tapia Stoll N.; Jaganathan S.; Natarajan U.; Rathinavelu A. Abstract 445: Epigenetic modification induced by DNMTs inhibitor enhances apoptosis through p21 pathway in neuroblastoma cells. Cancer Research 2025, 85(8_Supplement_1), 445–445;
DOI: 10.1158/1538-7445.AM2025-445.
Jaganathan S. S.; Natarajan U.; Rathinavelu A. Abstract 1618: Blocking MDM2 induces downregulation of PARP and enhances apoptosis in SK-N-SH neuroblastoma cells. Cancer
Research 2025, 85(8_Supplement_1), 1618–1618;
DOI: 10.1158/1538-7445.AM2025-1618.
Sobhan, M; Islam, M.M.; Mondal, A.M. “Interpreting Lung Cancer Health Disparity between African American Males and European American Males,” 2024 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Lisbon, Portugal, 2024, pp. 7141-7143,
DOI: 10.1109/BIBM62325.2024.10822014.
Ocasio, B. A.; Hu, J.; Stathias, V.; Martinez, M. J.; Burnstein, K. L.; Schurer, S. C. 3O Pre-clinical pan-cancer drug repurposing via deep learning. ESMO Open 2024, 9.
DOI: 10.1016/j.esmoop.2024.103749.
Pescov, J. G.; Martinez, M. J.; Schurer, S. 82P Applying computational approaches to build a predictive protein structure and discover novel inhibitors for mitotic serine/threonine kinase BUB1B. ESMO Open 2024, 9.
